First publication on chacma baboons submitted! Land use influence on chacma baboon (Papio ursinus) diet in South Africa using stable isotopes
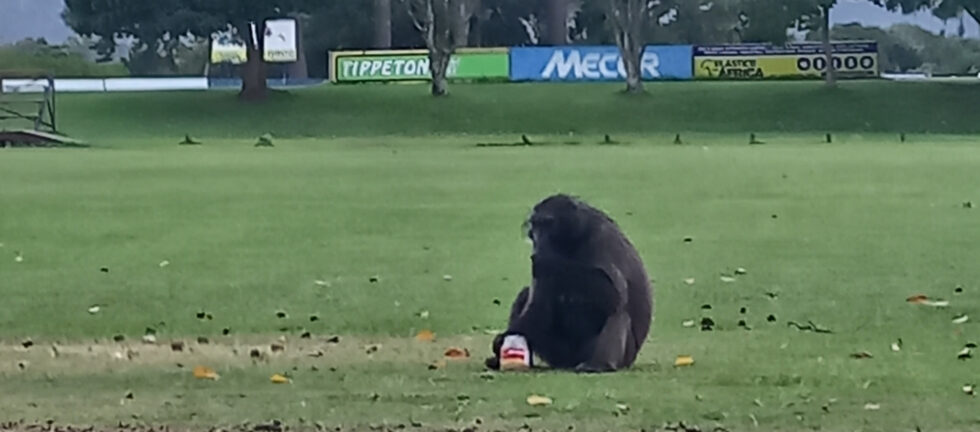
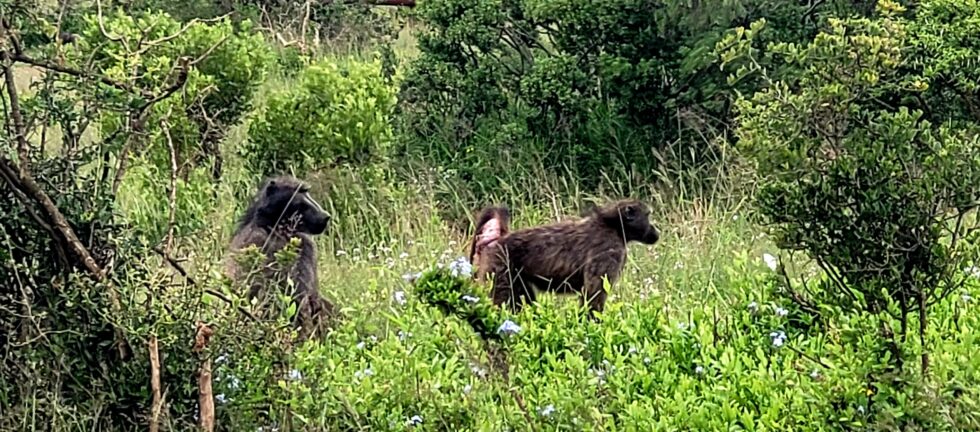
Anthropization processes affect wildlife feeding behaviours due to changes in resource availability related to land use and land cover change. To better understand the ecological responses of wildlife towards anthropogenic change, it is essential to evaluate whether human land use, characterized by high human-modified food availability, has an impact on wild animal feeding ecology. The chacma baboon (Papio ursinus) is interesting to study potential diet changes as it is largely present along a gradient of anthropized areas in Southern Africa. In this study, fecal samples from chacma baboon troops were collected in different land use habitats (peri-urban, agricultural and natural forest habitat) in the Garden Route, South Africa, and their isotopic ratios of carbon (δ13C) and nitrogen (δ15N) measured. Results showed significant differences between δ15N ratios according to land use, indicating significant higher protein intake in areas with human influence in comparison to natural forest habitats. Furthermore, the large majority of the collected samples were contained within the bracket that reflect the C3 ecosystem of the Garden Route region, with the exception of some samples showing higher δ13C ratios associated with the consumption of anthropogenic foods (containing sugar, corn and wheat). The potential protein increase, as well as sources of C4 plants present in the diets in anthropized areas suggests a visible dietary shift for this species between natural and transformed landscapes. In the future, it will be essential to determine whether and how the consumption of human-modified food could affect the health and associated fitness of chacma baboons.