A population genetic perspective on the origin, spread and adaptation of the human malaria agents Plasmodium falciparum and Plasmodium vivax

Abstract
Malaria is considered one of the most important scourges that humanity has faced during its history, being responsible every year for numerous deaths worldwide. The disease is caused by protozoan parasites, among which two species are responsible of the majority of the burden, Plasmodium falciparum and Plasmodium vivax. For these two parasite species, the questions of their origin (how and when they appeared in humans), of their spread throughout the world, as well as how they have adapted to humans have long been of interest to the scientific community. Here, we review the current knowledge that has accumulated on these different questions, thanks in particular to the analysis of the genetic and genomic variability of these parasites and comparison with related Plasmodium species infecting other host species (like non-human primates). In this paper we review the existing body of knowledge, including current research dealing with these questions, focusing particularly on genetic analysis and genomic variability of these parasites and comparison with related Plasmodium species infecting other species of host (such as non-human primates).
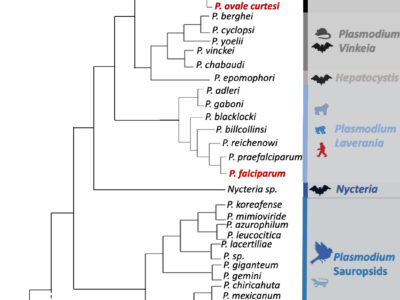
Fig. 1 Haemosporidian parasite schematic phylogenetic tree. The tree was drawn mostly based on the results of Galen et al. (2018). The six human malarial species are indicated in bold red. Colors indicate the different Plasmodium subgenus. Vertebrate host groups are represented by silhouette icons. Nodes that are not robust (i.e. varying positions of the branches depending on the study, gene used, species used for the rooting, etc.) are indicated with a black round in the phylogeny. For P. carteri, the branch has been represented with a dashed line because the species status and its position in the phylogeny need to be validated.
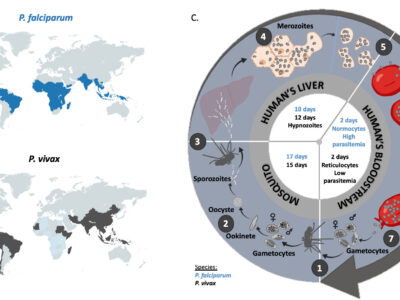
Fig. 2 Geographic distribution and life cycle comparison between P. falciparum and P. vivax. A. Map representation of the geographic incidence of P. falciparum, based on data from Weiss et al. (2019). B. Map representation of the geographic incidence of P. vivax, based on data from Battle et al. (2019) for high transmission regions and on data from Twohig et al. (2019) for low transmission regions in Africa (i.e. reports of all P. vivax occurrences in Africa). C. 1. A female anopheles mosquito ingests during its blood meal male and female gametocytes that mature rapidly into gametes. Inside the midgut of the mosquito, the male fertilizes the female gametocyte which results in the formation of a zygote. After meiosis, the resulting ookinete penetrates the mosquito gut wall. 2. The ookinete forms the oocyst, inside which the asexual replication will occur to give rise to several thousand sporozoites (sporogony). Once the oocyst is ruptured, sporozoites migrate to the salivary glands of the mosquito. 3. Malaria infection occurs once a female anopheles mosquito injects sporozoites inside the bloodstream of the vertebrate host (here a human) during a blood meal. 4. Injected sporozoites then reach liver, invade the hepatocytes and asexually multiply. 5. After 10 days for P. falciparum and 12 days for P. vivax, merozoites are released into the bloodstream. 6. The merozoites invade the erythrocytes (red blood cells; normocytes for P. falciparum and reticulocytes for P. vivax) and asexually multiply until the red cell burst. The released parasites infect other red blood cells and this cycle is repeated several times, causing fever each time parasites break free and invade blood cells. 7. At some point, some of the parasites may leave this asexual replication cycle and differentiate into male and female gametocytes which will eventually finish their course into an anopheles mosquito gut during a blood meal. P. falciparum and P. vivax biological characteristics are indicated in blue and black respectively inside the cycle.
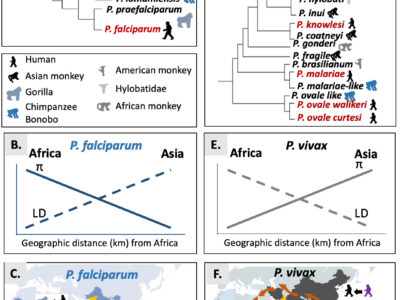
Fig. 3 Origin and spread of P. falciparum and P. vivax. A. Schematic phylogenetic relationships representations in the Laverania. B. Observed trends for the genetic diversity (P) and linkage disequilibrium (LD) for P. falciparum. C. Suggested scenario for the African origin and potential routes of migration for P. falciparum. The original source is indicated by the red point: Africa for P. falciparum. Yellow arrows represent the suggested dispersal routes of P. falciparum from Central Africa. D. Schematic phylogenetic relationships representations in the Plasmodium subgenus. E. Observed trends for the genetic diversity (P) and linkage disequilibrium (LD) for P. vivax. F. Suggested scenarios for the African or Asian origin and potential routes of migration of P. vivax. The original sources are indicated by a red point when considered in Africa and a green point when considered in Asia. Yellow arrows represent the suggested dispersal routes of P. vivax from Central Africa. Brown arrows represent the suggested dispersal routes of P. vivax when its origin is considered in Asia.
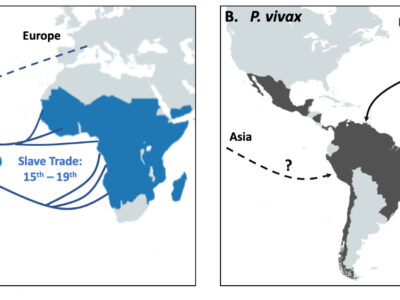
Fig. 4 Schematic representation of the proposed origin in America of P. falciparum in panel A and P. vivax in panel B. Main routes of the transatlantic slave trades, major European Empires between the 15th and 19th centuries in America are indicated with arrows. When the migration route and origin have been confirmed by genetic studies, arrows indicate the direction of the transfer. When migrations routes are still under debate, dashed arrows indicate the sense of the suggested transfer.
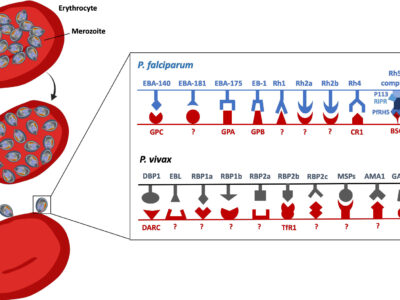
Fig. 5 Schematic representation of some of the main parasite ligand – host receptors interactions involved in erythrocyte invasion for P. falciparum and P. vivax. At the end of the erythrocytic cycle, schizonts burst and release merozoites in the blood stream, allowing invasion of new uninfected red blood cells. The recognition of red blood cells and the invasion process require specific interactions between the parasite ligands and the red blood cells receptors of the vertebrate host. The invasion process might occur through complete unknown pathways with both unidentified ligands (noted with a ?) and receptors (also noted ?). For P. falciparum and P. vivax some parasite ligands are pseudogeneised (nonfunctional): these ligands have been indicated in the figure with dashed structures. EBA-140, erythrocyte binding antigen 140; EBA-181, erythrocyte binding antigen 181; EBA-175, erythrocyte binding antigen 175; EBA165, erythrocyte binding antigen 165; EB1, erythrocyte binding like protein 1; RH1, Reticulocyte binding-like protein 1; RH2a, Reticulocyte binding-like protein 2a; RH2b, Reticulocyte binding-like protein 2b; RH3, Reticulocyte binding-like protein 3; RH4, Reticulocyte binding-like protein 4; RH5 complex, Reticulocyte binding-like protein 5 complex; GPC, glycophporin C; GPA, glycophorin A; GPB, glycophorin B; CRP1, complement receptor 1; BSG, Basigin; AMA1, anchored micronemal antigen 1; DBP, Duffy binding protein; EBP, erythrocyte-binding protein; GAMA, glycosylphosphatidylinositol-anchored micronemal antigen; MSP, merozoite surface protein; RBP, reticulocyte binding proteins; DARC; Duffy antigen receptor for chemokines; TfR1, transferrin receptor 1.

Publication infos
- Virginie Rougeron, Larson Boundenga, Céline Arnathau, Patrick Durand, François Renaud, Franck Prugnolle
- virginie.rougeron@cnrs.fr and prugnolle@gmail.com
- Laboratory MIVEGEC, University of Montpellier, CNRS, IRD, 900 rue Jean François Breton, 34090 Montpellier, France. - CREES, Centre de Recherches en Écologie et Évolution de la Santé, Montpellier, France. - CIRMF, Centre Interdisciplinaire de Recherches Médicales de Franceville, Franceville, Gabon.
- This work was supported by the ANR t-ERC EVAD 2017, ANR JCJC GENAD 2021, ANR MICETRAL 2020, the labex CEMEB (project CONVERGE) and the PEPS ECOMOB project MOV.
See the publication
FEMS Microbiology Reviews, fuab047
DOI : 10.1093/femsre/fuab047